Huntington's Disease and C. elegans - Michael Babbitt, Isaac Chrisman, Annie Kelly, Gian Lagemann, Max Lee, Eric Ramirez - Grand Challenges Initiative

Sensors | Free Full-Text | Neural Network-Based Autonomous Search Model with Undulatory Locomotion Inspired by Caenorhabditis Elegans

Natural genetic variation drives microbiome selection in the Caenorhabditis elegans gut - ScienceDirect

Bounded rationality in C. elegans is explained by circuit-specific normalization in chemosensory pathways | Nature Communications
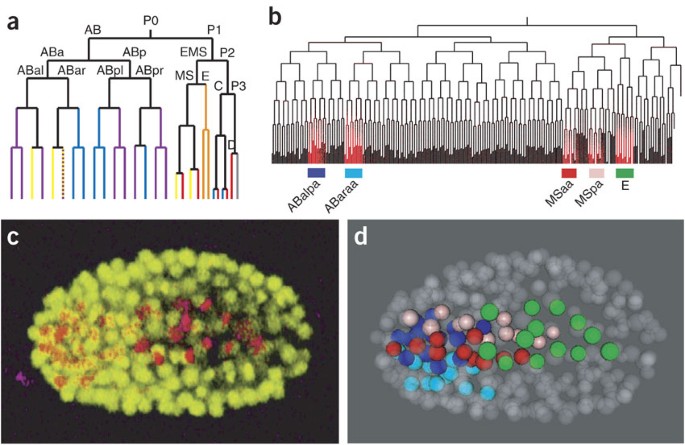
Automated analysis of embryonic gene expression with cellular resolution in C. elegans | Nature Methods
Metabolomic signature associated with reproduction-regulated aging in Caenorhabditis elegans - Figure f2 | Aging

Systems‐level quantification of division timing reveals a common genetic architecture controlling asynchrony and fate asymmetry | Molecular Systems Biology
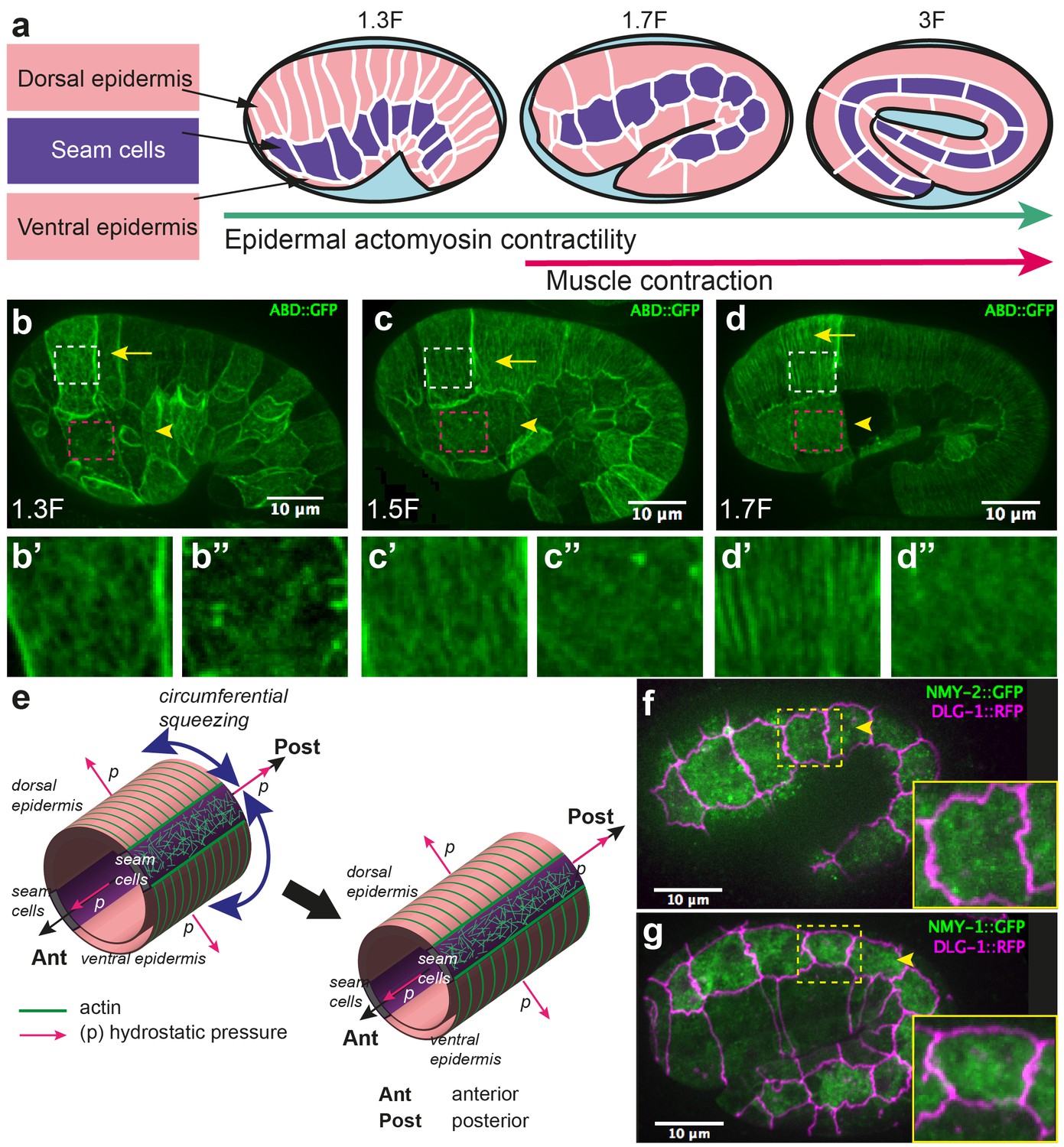
Figures and data in The interplay of stiffness and force anisotropies drives embryo elongation | eLife
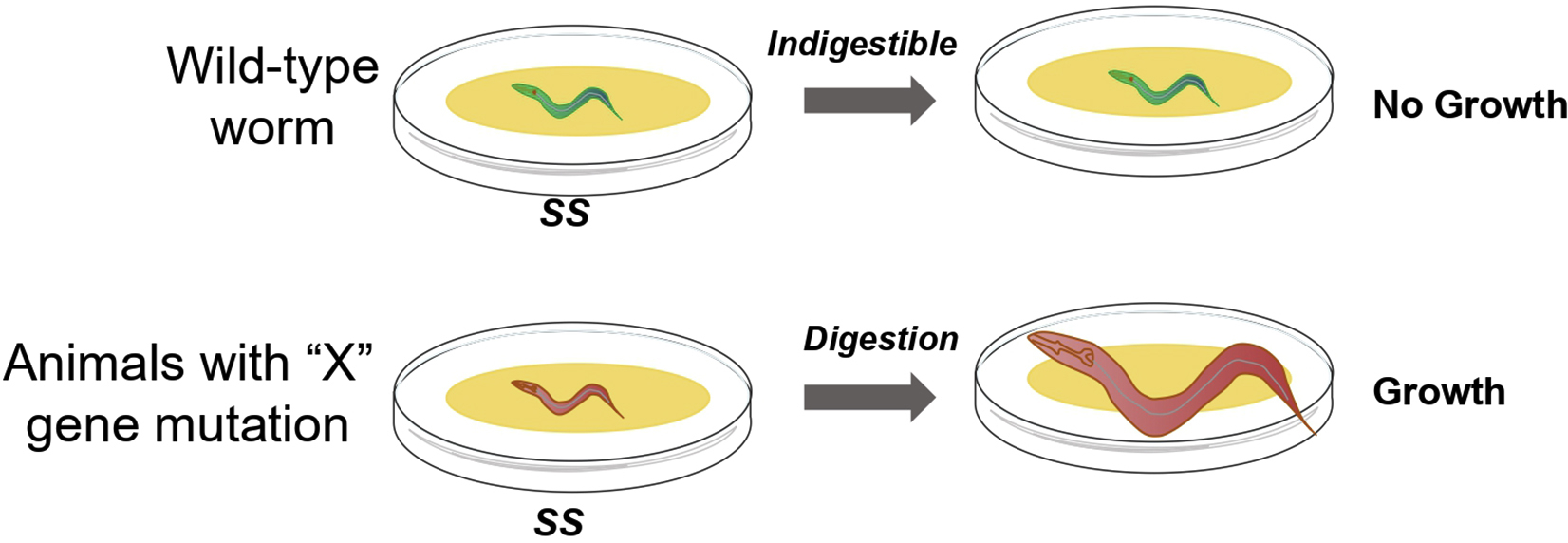
Protocol for investigating the effect of food digestion in C. elegans on development by feeding the inedible bacteria Staphylococcus saprophyticus